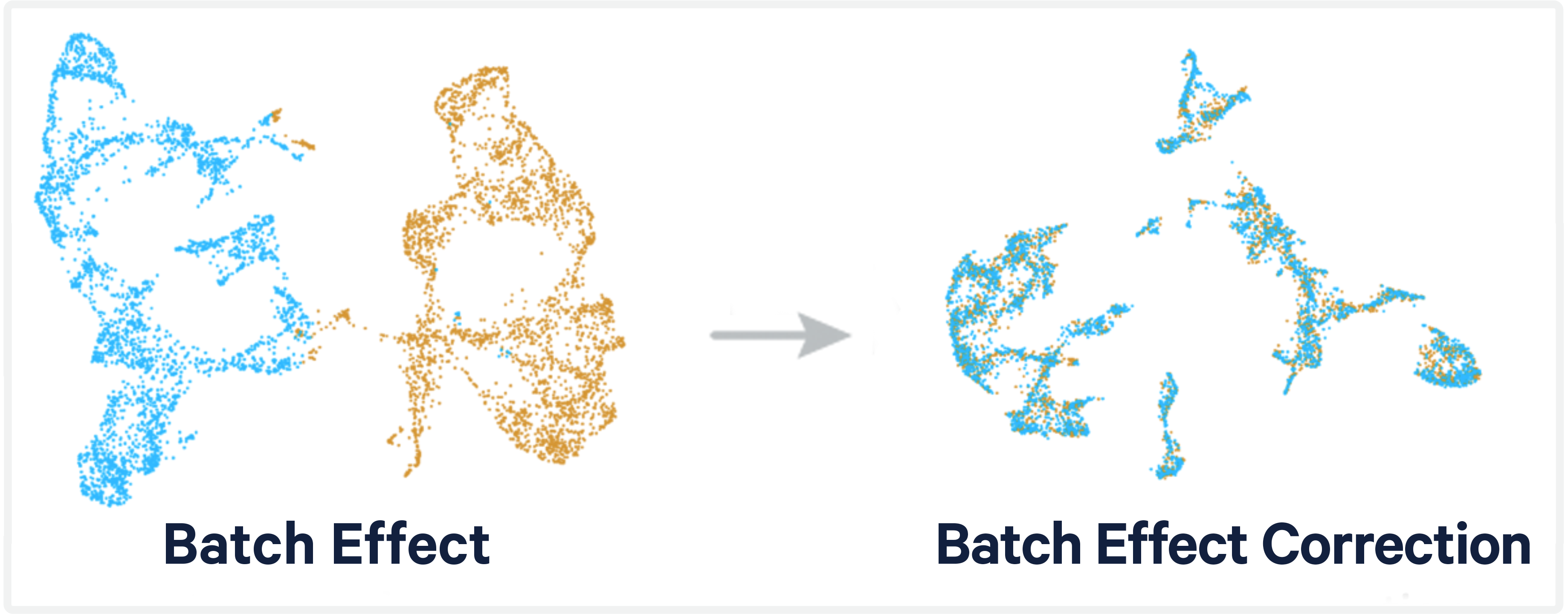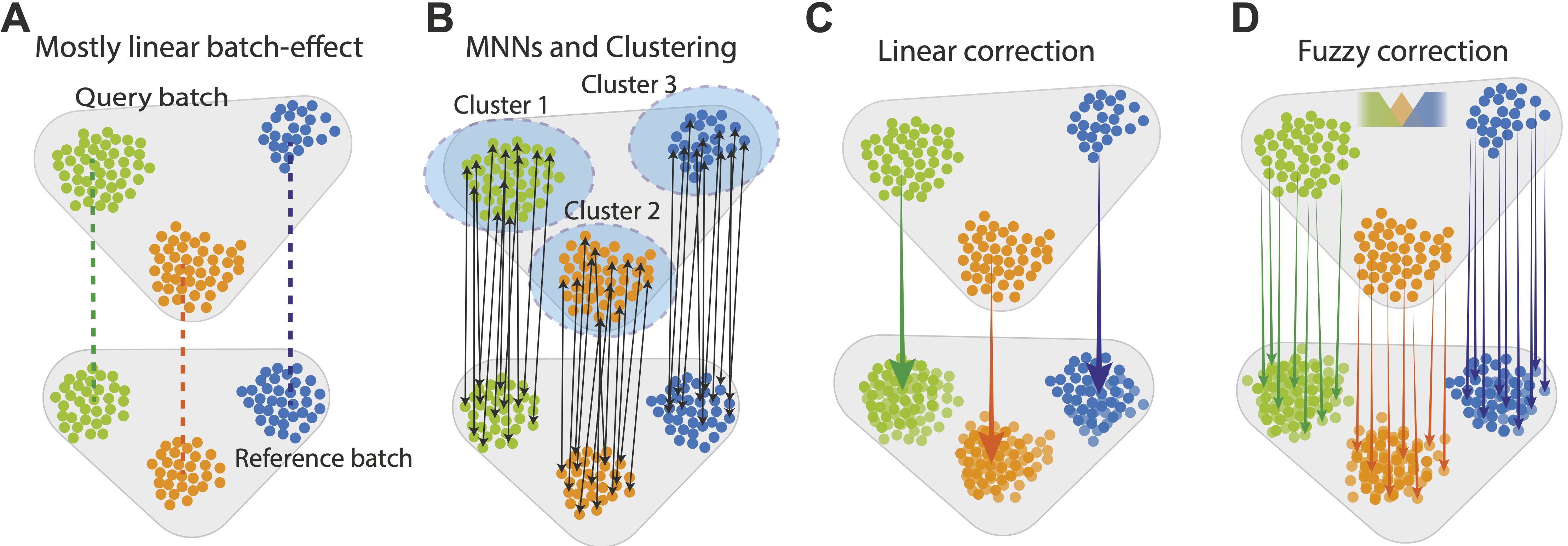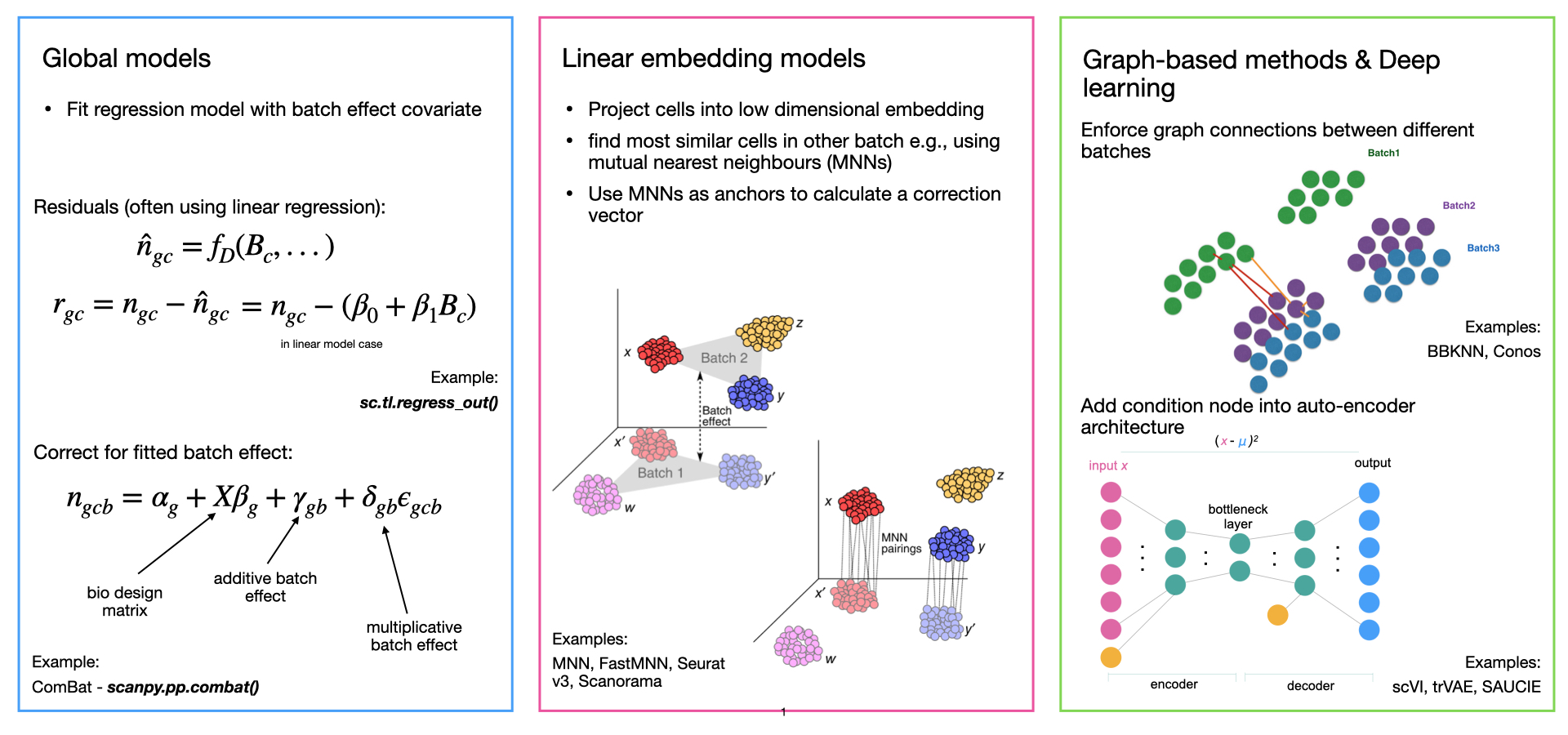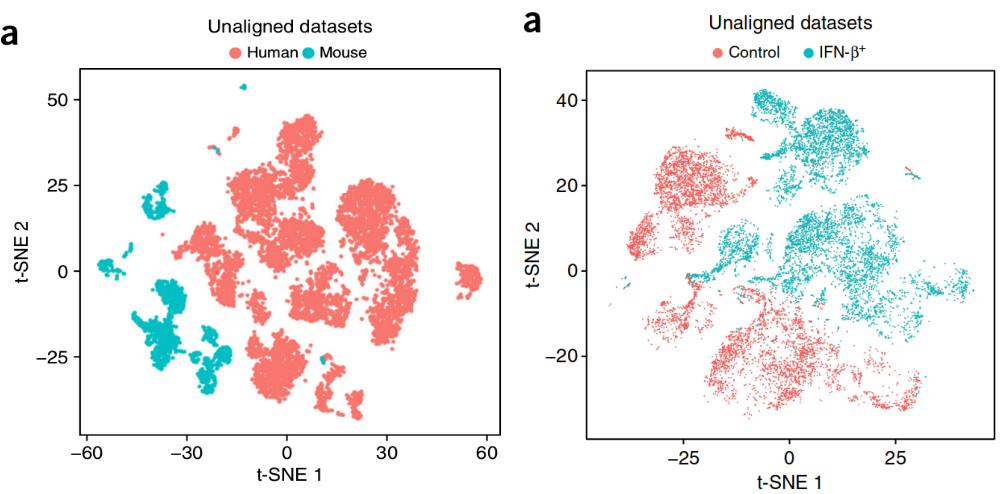
How to Batch Correct Single Cell. Comparing batch correction methods for… | by Nikolay Oskolkov | Towards Data Science

BatchServer: A Web Server for Batch Effect Evaluation, Visualization, and Correction | Journal of Proteome Research
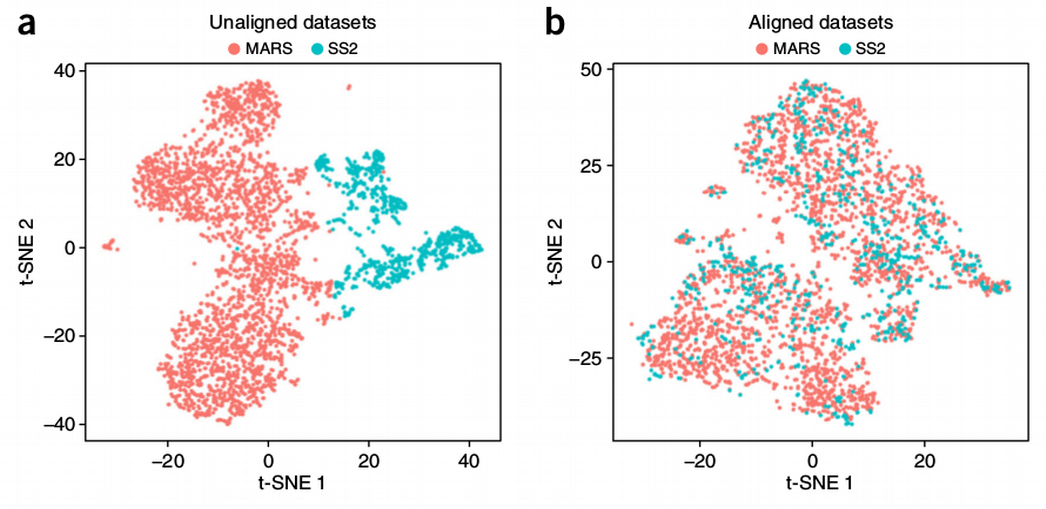
How to Batch Correct Single Cell. Comparing batch correction methods for… | by Nikolay Oskolkov | Towards Data Science
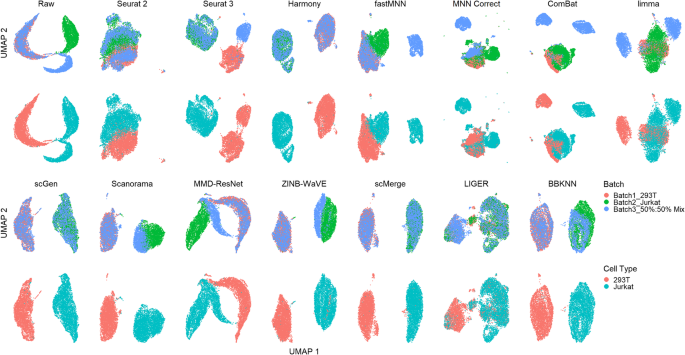
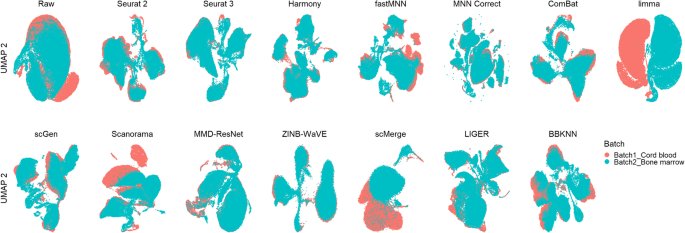
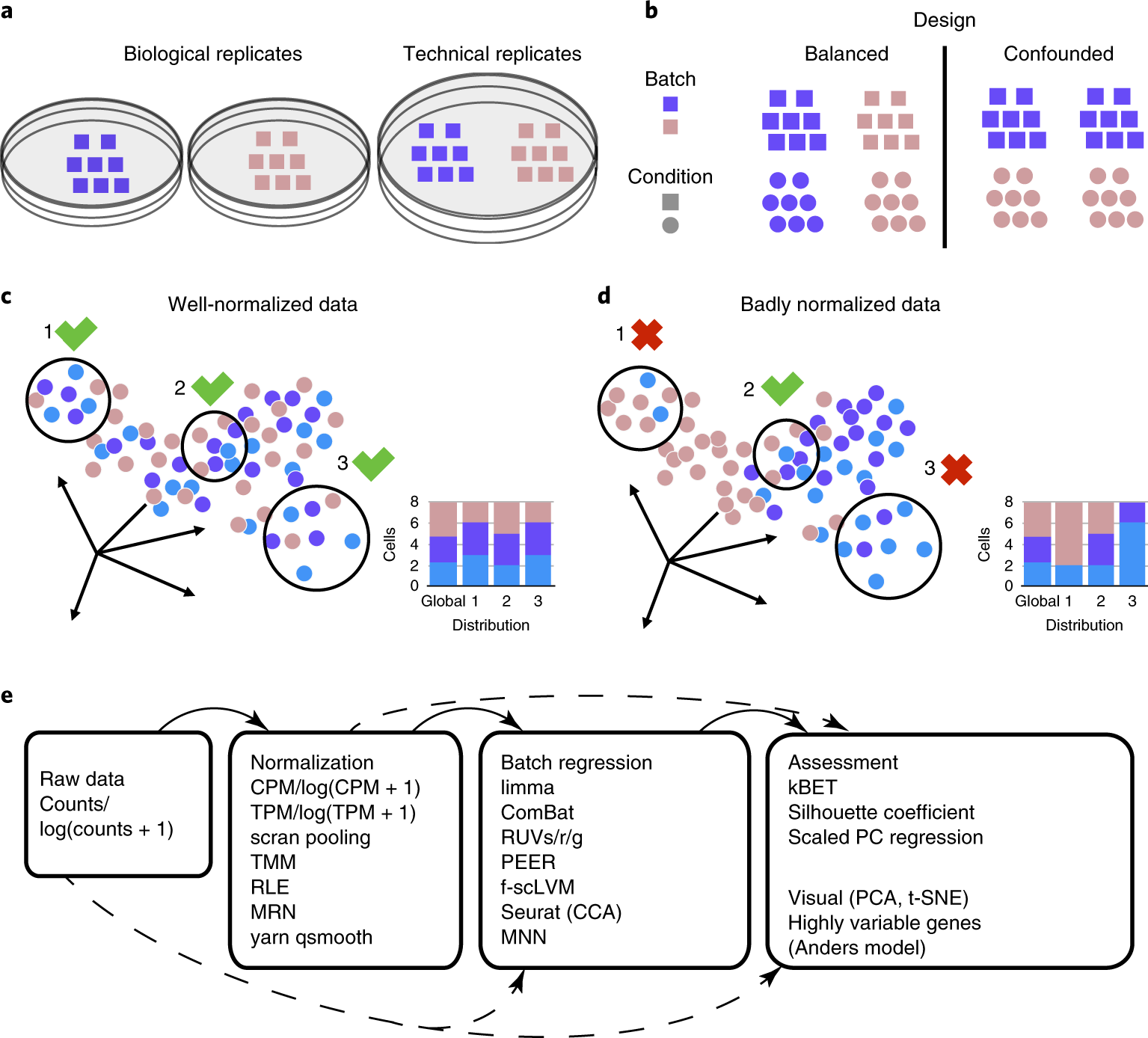
A benchmark of batch-effect correction methods for single-cell RNA sequencing data | Genome Biology | Full Text
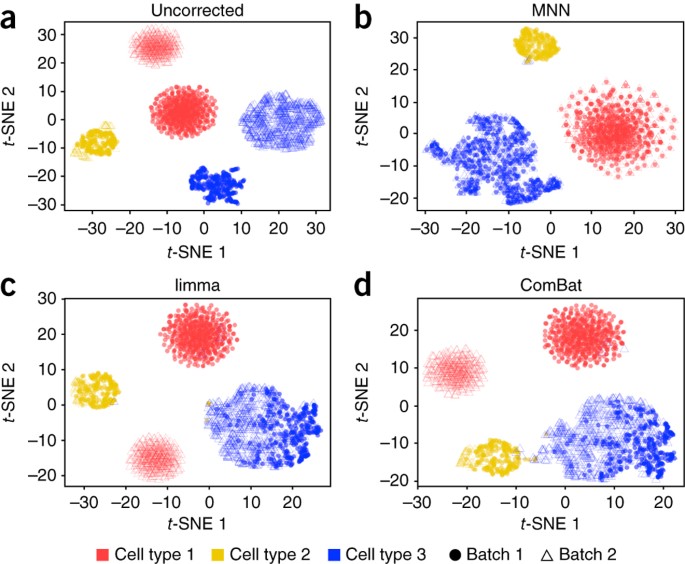
Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors | Nature Biotechnology

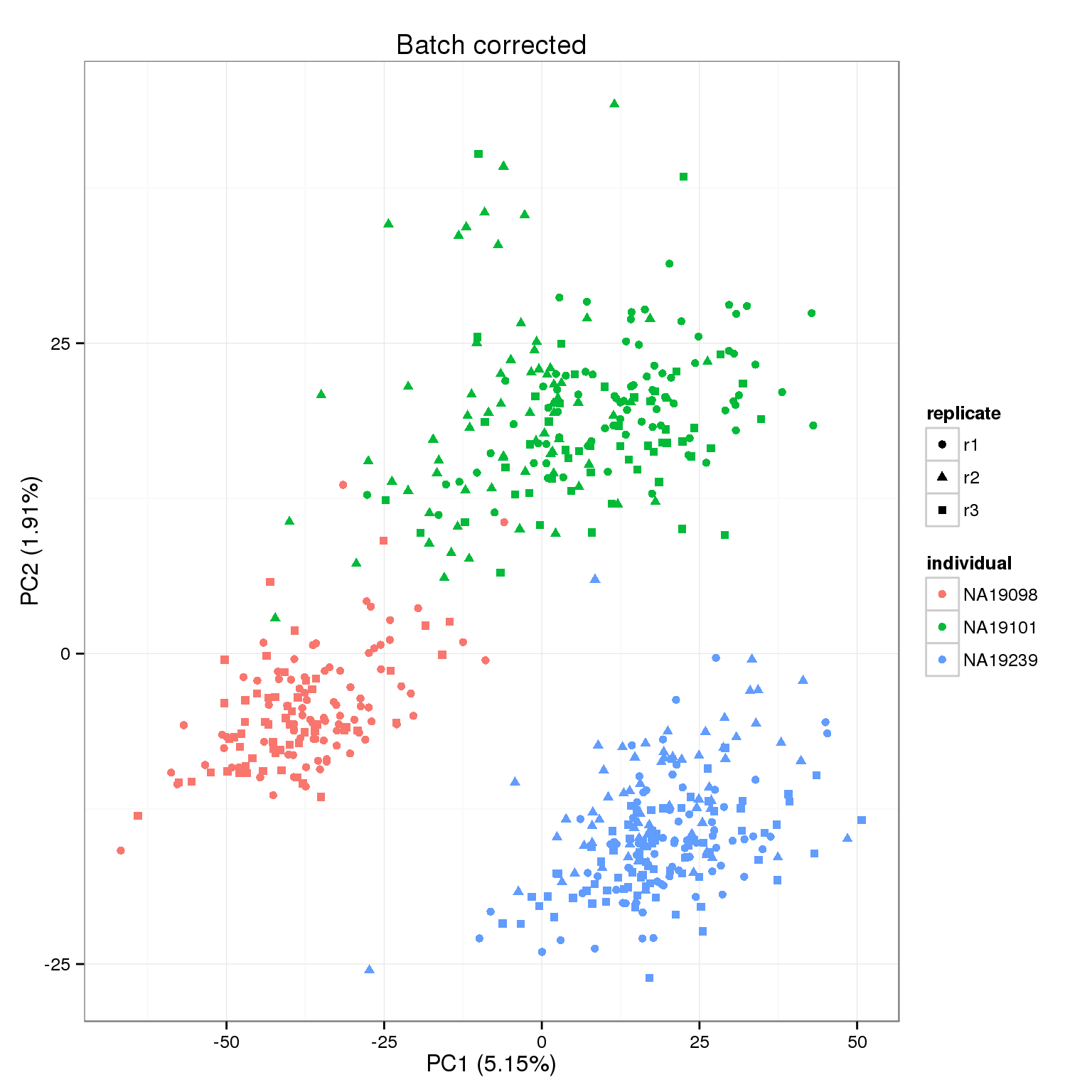
Batch effect correction of SMART-seq2 dataset. (A) Batch effect was... | Download Scientific Diagram
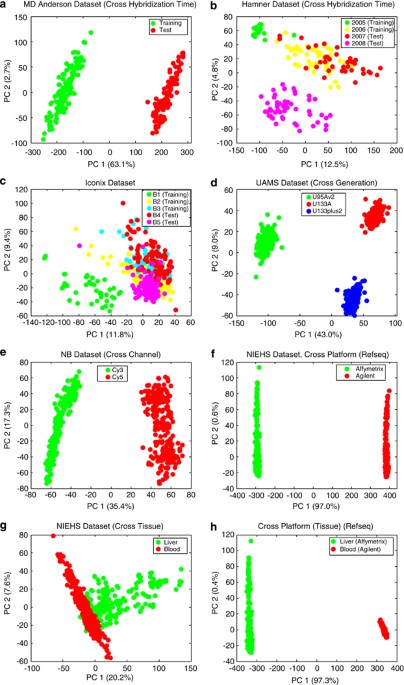
A comparison of batch effect removal methods for enhancement of prediction performance using MAQC-II microarray gene expression data | The Pharmacogenomics Journal

Perspectives for better batch effect correction in mass-spectrometry-based proteomics - Computational and Structural Biotechnology Journal
Removing Batch Effects from Longitudinal Gene Expression - Quantile Normalization Plus ComBat as Best Approach for Microarray Transcriptome Data | PLOS ONE

rna seq - Batch effect correction using a subset of samples - using DE genes - Bioinformatics Stack Exchange

Correction of batch effect in CRC and normal gene expression datasets... | Download Scientific Diagram

Scaling Method for Batch Effect Correction of Gene Expression Data Based on Spectral Clustering | Bentham Science
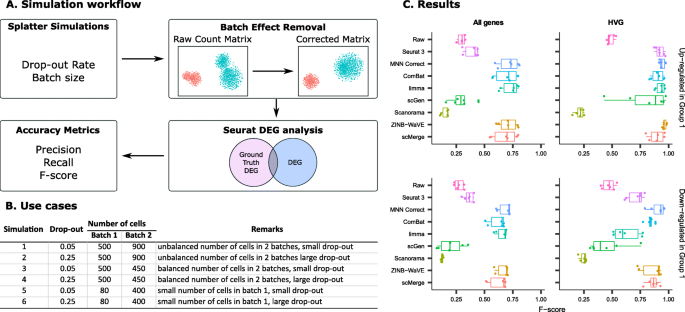
A benchmark of batch-effect correction methods for single-cell RNA sequencing data | Genome Biology | Full Text

Batch correction methods for nontarget chemical analysis data: application to a municipal wastewater collection system | Analytical and Bioanalytical Chemistry

Correcting batch effects in single-cell RNA sequencing data by matching mutual nearest neighbours | bioRxiv
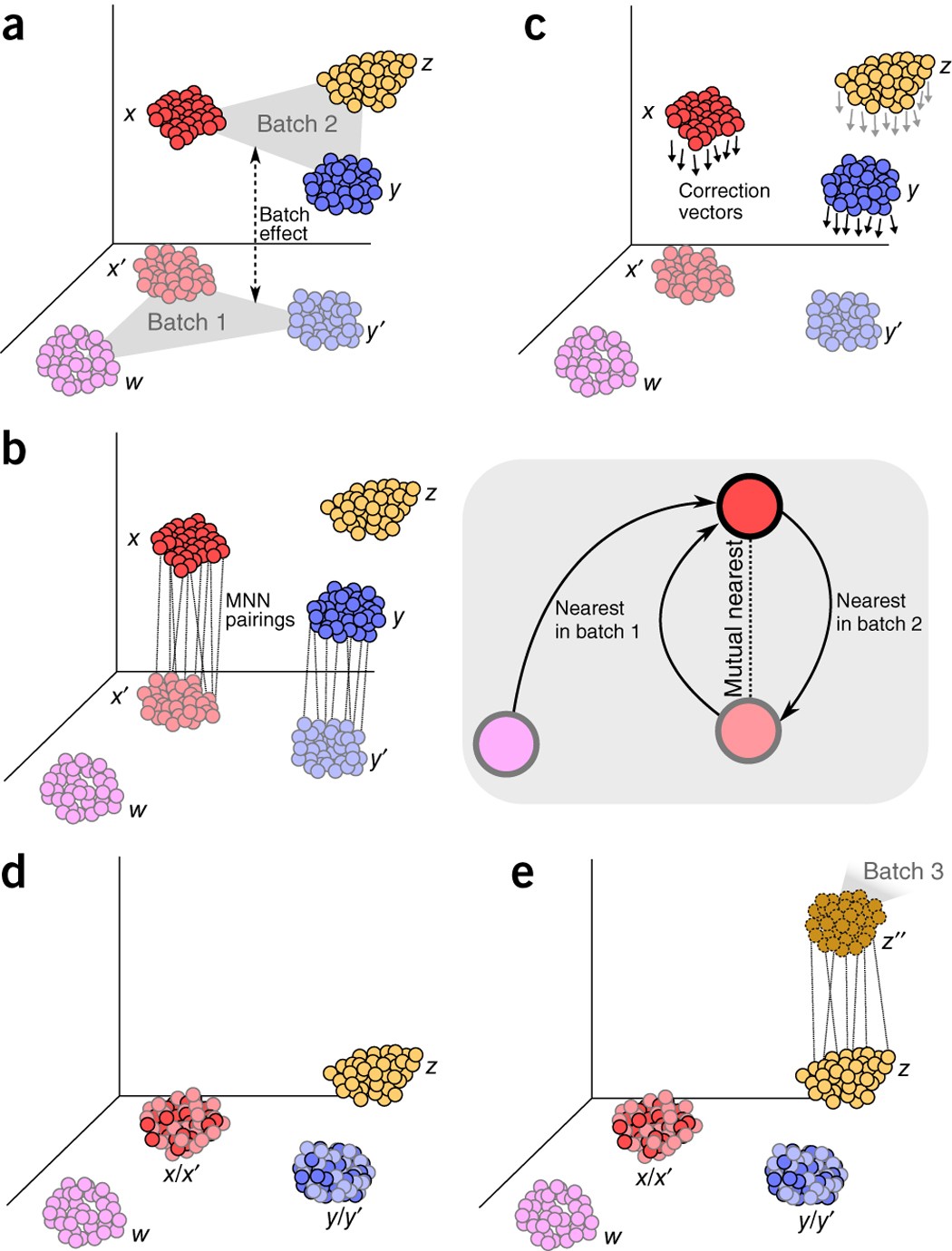
Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors | Nature Biotechnology